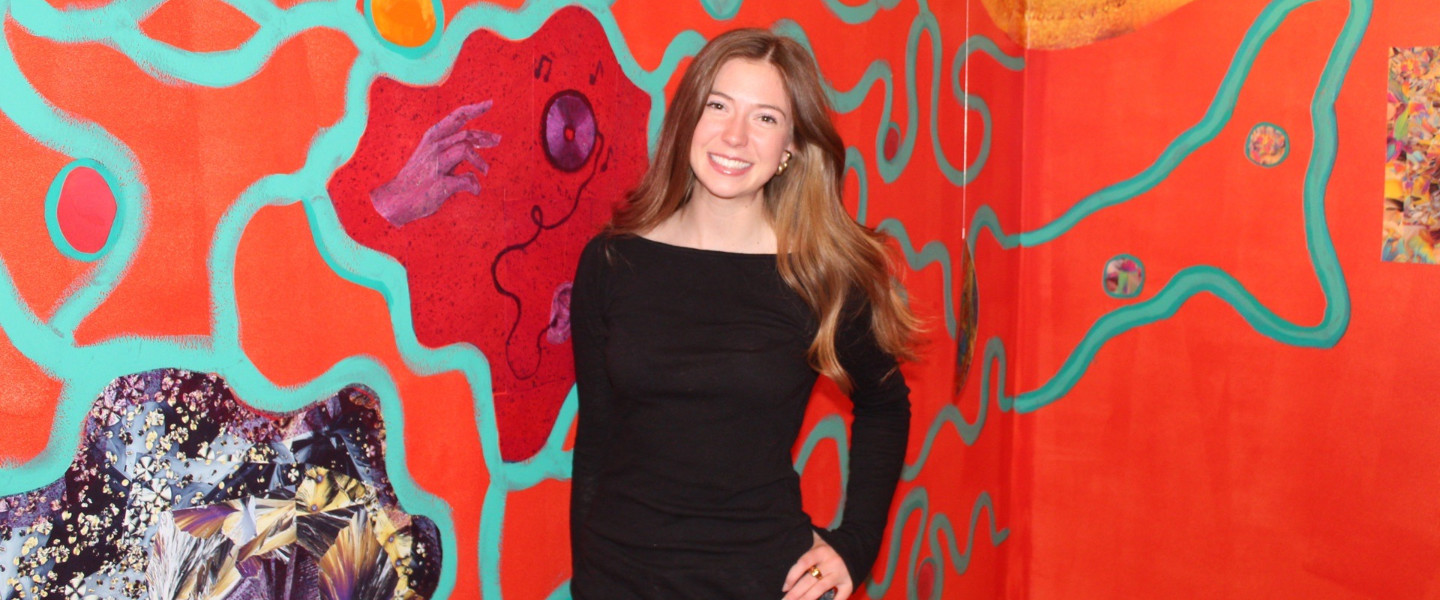
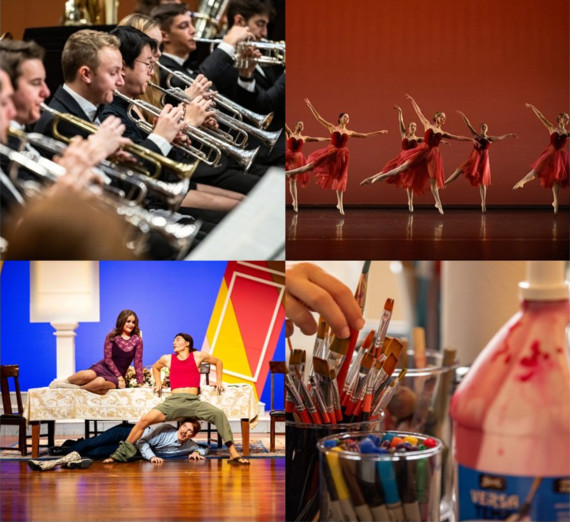


ARTS at GONZAGA
Celebrate music, theater, visual arts, and more—experience the vibrant creativity of our students and faculty.
Empowering individuals to thrive in intellect, vocation, and spirit, equipping them to embrace the challenges of today and tomorrow with resilience, purpose and a commitment to serving others.
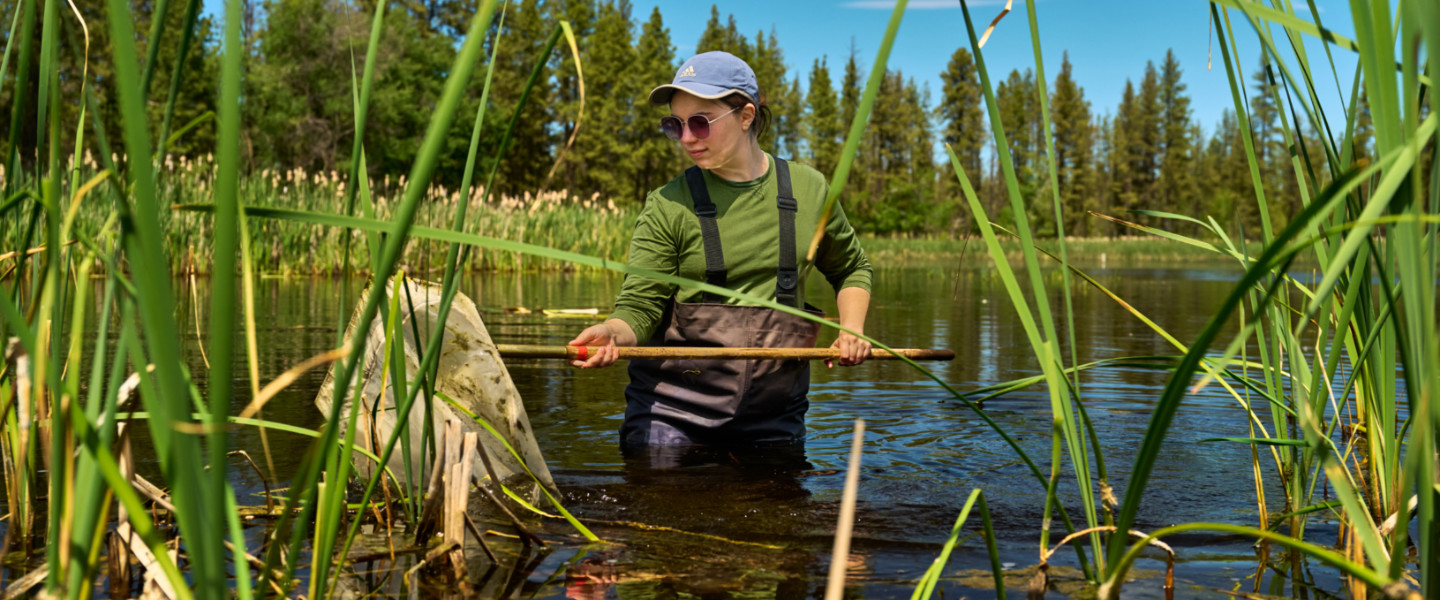
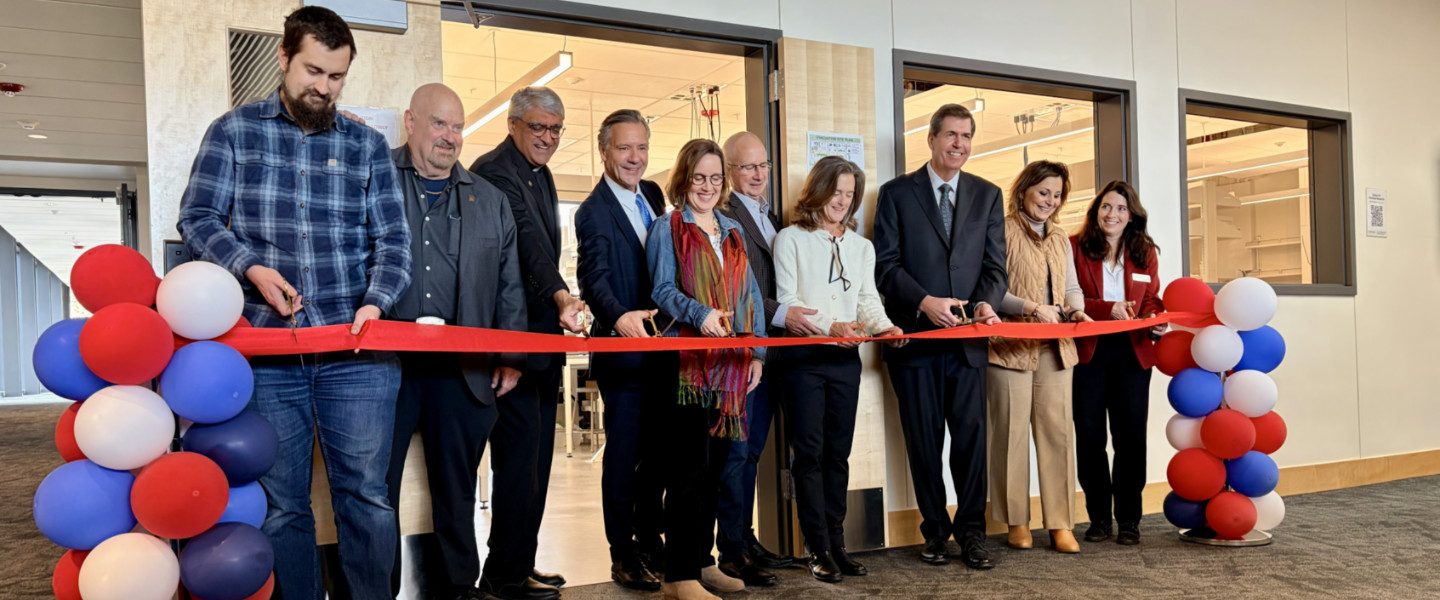




Celebrate music, theater, visual arts, and more—experience the vibrant creativity of our students and faculty.
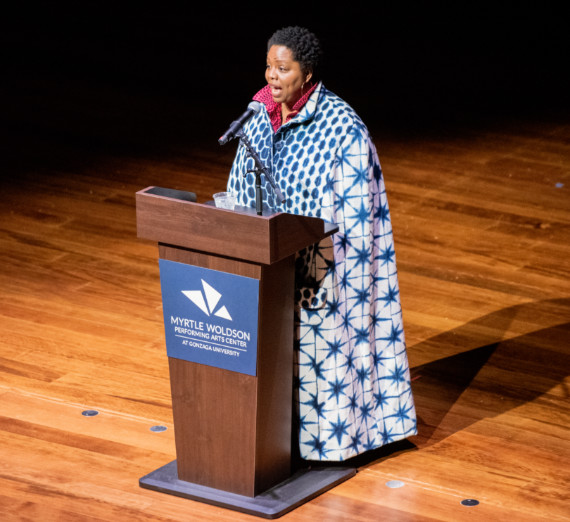

Engage with distinguished speakers and thought-provoking conversations that inspire learning and dialogue.
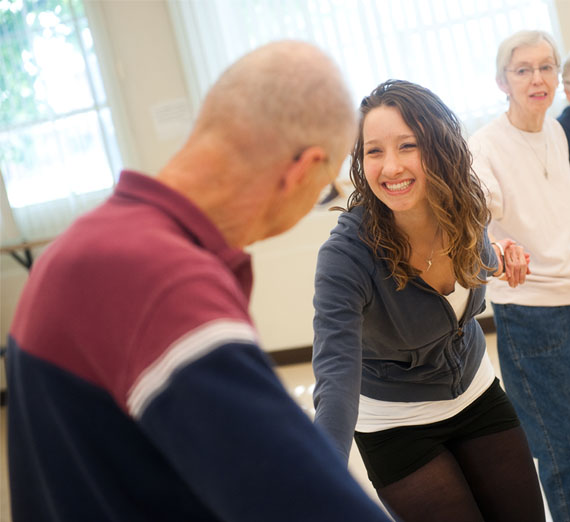

See how Gonzaga collaborates with organizations and communities to create meaningful impact locally and globally.
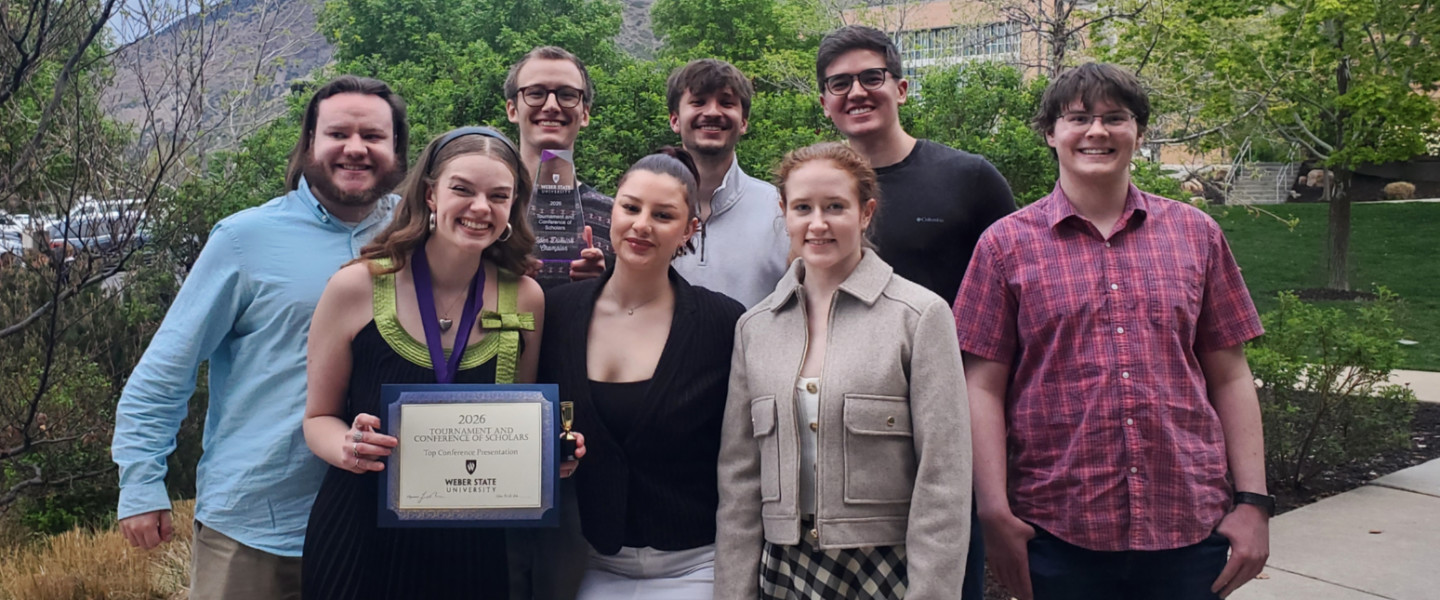
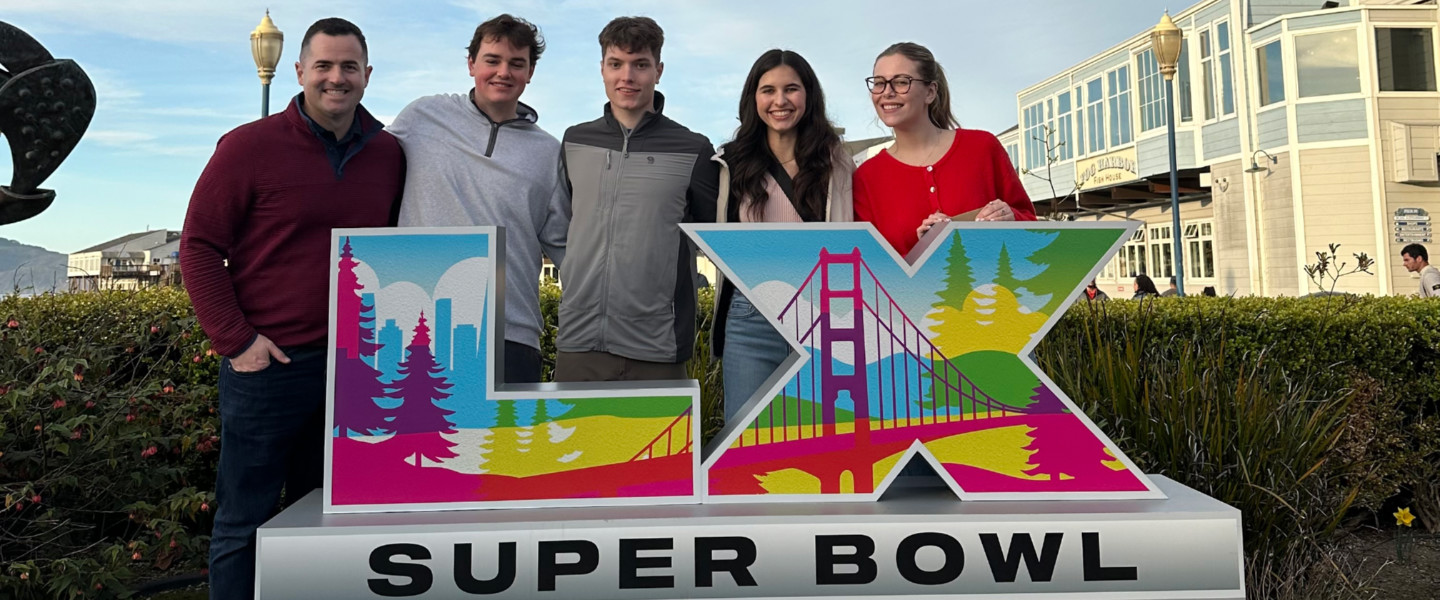
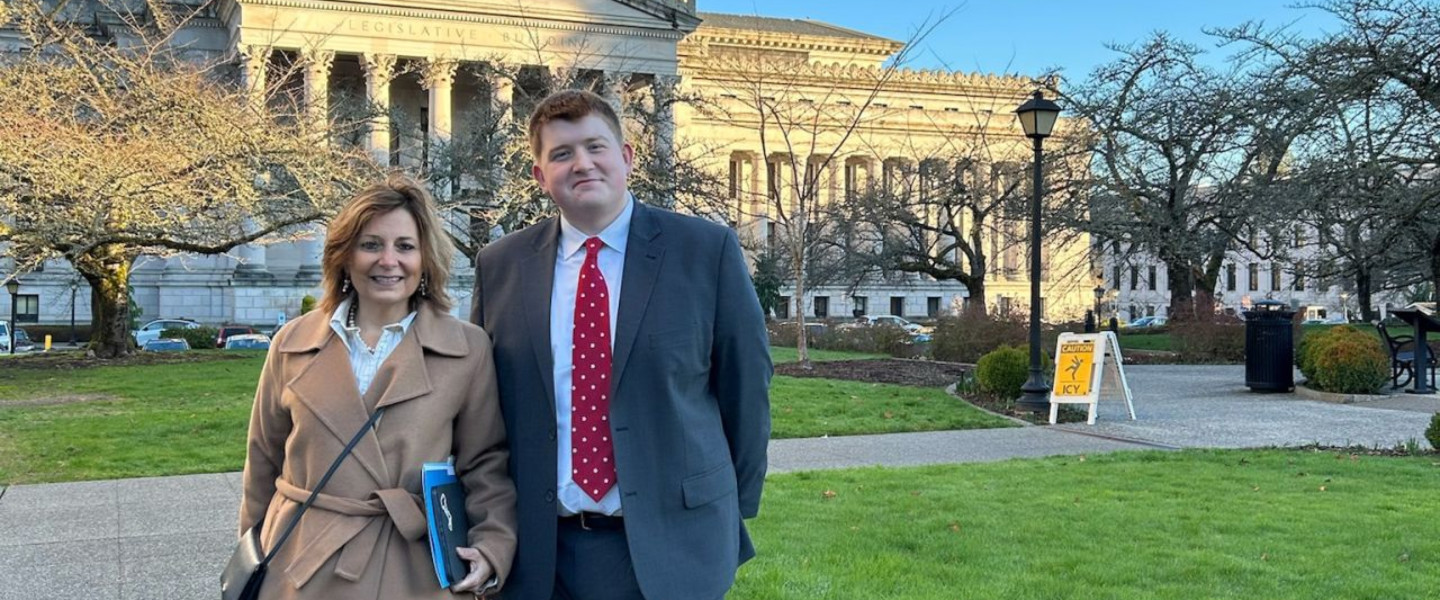



Gonzaga University students can earn credit working on real projects with Intel, Netflix and Airbnb...
Gonzaga alumni who work at Microsoft open doors for students
Gonzaga University debate team wins its third consecutive CARD National Championship. Senior Kaelyn...
The College of Arts and Sciences at Gonzaga is supported by the generosity of Zags just like you. Thanks to donations from families, alumni, faculty, staff, friends and more, the College is empowered to provide a distinctive, Jesuit, liberal arts education that inspires graduates to pursue a better world.